
The outreach program will develop infrastructure and methods to involve a large number of researchers, faculty, and students in the analysis of the maize genome. During the outreach program we will focus on training researchers and faculty members on how to use workflows and bioinformatics programs to analyze the expected MaizeCODE data sets. Researchers, faculty, and students will be poised to participate in de novo explorations of the structure and function of the maize genome.
2020 Publication
Frontiers in Plant Science
Management, Analyses, and Distribution of the MaizeCODE Data on the Cloud
Liya Wang¹, Zhenyuan Lu¹, Melissa delaBastide¹, Peter Van Buren¹, Xiaofei Wang¹, Cornel Ghiban¹, Michael Regulski¹, Jorg Drenkow¹, Xiaosa Xu¹, Carlos Ortiz-Ramirez2 , Cristina F.
Marco¹, Sara Goodwin¹, Alexander Dobin¹, Kenneth D. Birnbaum²,
David P. Jackson¹, Robert A. Martienssen¹, William R. McCombie¹, David A. Micklos¹, Michael C.
Schatz¹³, Doreen H. Ware¹⁴* and Thomas R. Gingeras¹*
¹ Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, United States, ² New York University, New
York, NY, United States, ³ Johns Hopkins University, Baltimore, MD, United States, ⁴ USDA-ARS Robert W. Holley Center for
Agriculture and Health, Ithaca, NY, United States
Frontiers in Plant Science, published: March 31, 2020 | https://doi.org/10.3389/fpls.2020.00289
Abstract:
MaizeCODE is a project aimed at identifying and analyzing functional elements in the maize genome. In its initial phase, MaizeCODE assayed up to five tissues from four maize strains (B73, NC350, W22, TIL11) by RNA-Seq, Chip-Seq, RAMPAGE, and small RNA sequencing. To facilitate reproducible science and provide both human and machine access to the MaizeCODE data, we enhanced SciApps, a cloud-based portal, for analysis and distribution of both raw data and analysis results. Based on the SciApps workflow platform, we generated new components to support the complete cycle of MaizeCODE data management. These include publicly accessible scientific workflows for the reproducible and shareable analysis of various functional data, a RESTful API for batch processing and distribution of data and metadata, a searchable data page that lists each MaizeCODE experiment as a reproducible workflow, and integrated JBrowse genome browser tracks linked with workflows and metadata. The SciApps portal is a flexible platform that allows the integration of new analysis tools, workflows, and genomic data from multiple projects. Through metadata and a ready-to-compute cloud-based platform, the portal experience improves access to the MaizeCODE data and facilitates its analysis.
2019 Publication
PLoSONE
Double triage to identify poorly annotated genes in maize: The missing link in community curation
Marcela K. Tello-Ruiz, Cristina F. Marco, Fei-Man Hsu, Rajdeep S. Khangura, Pengfei Qiao, Sirjan Sapkota, Michelle C. Stitzer, Rachael Wasikowski, Hao Wu, Junpeng Zhan, Kapeel Chougule, Lindsay M. Barone, Cornel Ghiban, Demitri Muna, Andrew C. Olson, Liya C. Wang, Doreen C. Ware, David A. Micklos
PLoSONE 14(10): e0224086, published: October 28, 2019 https://doi.org/10.1371/journal.pone.0224086
Abstract:
The sophistication of gene prediction algorithms and the abundance of RNA-based evidence for the maize genome may suggest that manual curation of gene models is no longer necessary. However, quality metrics generated by the MAKER-P gene annotation pipeline identified 17,225 of 130,330 (13%) protein-coding transcripts in the B73 Reference Genome V4 gene set with models of low concordance to available biological evidence. Working with eight graduate students, we used the Apollo annotation editor to curate 86 transcript models flagged by quality metrics and a complimentary method using the Gramene gene tree visualizer. All of the triaged models had significant errors–including missing or extra exons, non-canonical splice sites, and incorrect UTRs. A correct transcript model existed for about 60% of genes (or transcripts) flagged by quality metrics; we attribute this to the convention of elevating the transcript with the longest coding sequence (CDS) to the canonical, or first, position. The remaining 40% of flagged genes resulted in novel annotations and represent a manual curation space of about 10% of the maize genome (~4,000 protein-coding genes). MAKER-P metrics have a specificity of 100%, and a sensitivity of 85%; the gene tree visualizer has a specificity of 100%. Together with the Apollo graphical editor, our double triage provides an infrastructure to support the community curation of eukaryotic genomes by scientists, students, and potentially even citizen scientists.
2020
Maize Annotation Jamboree
First Virtual Maize Annotation Jamboree, March 10-12, 2020
The 2020 Virtual Maize Annotation Jamboree (VMAJ) was organized by Cristina F. Marco (DNALC CSHL) and Marcela Karey Tello-Ruiz (Cold Spring Harbor Laboratory or CSHL) under the supervision of PI Doreen Ware (CSHL and USDA ARS).
The jamboree aims were: 1) identify and correct potential gene model errors in the draft B73 Zea mays V5 gene models, 2) train participants on the use of gene curation tools, and 3) establish a network of researchers and teaching faculty to support and implement genome annotation projects as Course-based Undergraduate Research Experiences (CUREs). Among the 28 remote participants were 20 of the original travel awardees (10 female, 10 male) from 13 higher education institutions including 4 Primarily Undergraduate Institutions (PUI), and the USDA ARS.
Zoom gallery view of participants on the final day of the VMAJ. From left to right and top to bottom and left to right: Arun Somwarpet-Seetharam (Iowa State University), Cristina F. Marco (DNALC-CSHL), Kevin R. Ahern (Cornell University), Brit Moss (Whitman College), Aman Kaur (Purdue University), Adrienne Kleintop (Delaware Valley University), Jianing Liu (University of Georgia), Marcela Karey Tello-Ruiz (CSHL), John Gray (The University of Toledo), Doreen Ware (CSHL), Elly Porestky (UC San Diego), Vincent Colantonio (University of Florida), Brian Zebosi (Iowa State University), Singha Dhungana (University Missouri Columbia), Vivek Shrestha (University Missouri Columbia), Nancy Manchada (Iowa State University), Raksha Singh (Purdue University), and Usha Bhatta (University of Georgia). Missing in the picture: Carrie Olson-Manning (Augustana University), Surinder Chopra (Penn State University), Kevin Begcy (University of Georgia), Ankita Abnave (The University of Toledo), Raksha Singh (Purdue University), Manwinder Singh Brar (Clemson University), and Devon Birdseye (UC San Diego).
During the course of the jamboree, participants were trained on the use of gene curation tools, whereby 686 genes were evaluated using the Gramene Gene Tree Visualizer and 42 genes reviewed in the Apollo B73+ gene editor. Over 95% of participants reported improved understanding of gene curation using the Gene Tree Visualizer and Apollo B73+ gene editor. As a follow up to the jamboree we have established monthly virtual meetings to share results, improve on the existing training material, and develop and support student projects and CUREs.) We are also continuing to host the maize annotation infrastructure (NAM Gene Tree Visualizer and Apollo B73+ server), weekly office hours, and a Slack instant messaging channel.
We believe this first maize virtual jamboree will serve as a proof-of-concept for similar future community curation efforts to improve genomic annotations in maize, and other important crops, and provides an example of opportunities for remote learning in a virtual and diverse collaborative environment.
The event was jointly supported through funding from NSF IOS-1127112, MCB-1744001, IOS-1445025 and USDA ARS 8062-21000-041.
2019
Maize Annotation Jamborees
January Jamboree for Faculty from PUIs
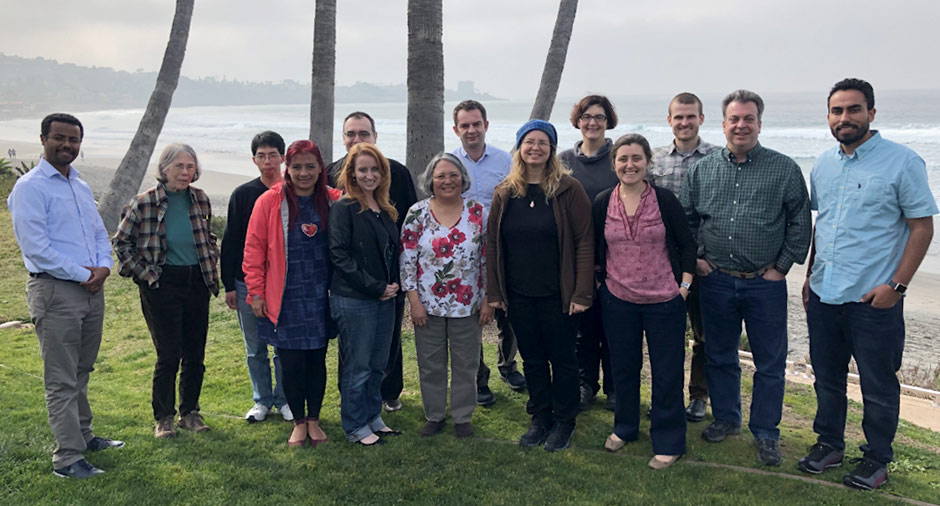
The second Maize Annotation Jamboree—the first designed for primarily undergraduate institution (PUI) faculty—was held January 10-11, 2019 at the Scripps Institution of Oceanography in San Diego, CA. The Jamboree was designed to train PUI faculty in the use of genome annotation tools as part of the community curation effort to improve the B73 Zea mays v4 gene models. The ultimate goal is to help faculty integrate maize annotation activities as course-based undergraduate research experiences (CUREs).
Eight applicants participated in this two-day event held adjacent to the annual International Plant and Animal Genome meeting, which half went on to attend.
This year’s participants were affiliated with eight institutions across the United States: Paul Bilinski (West Shore Community College), James Godde (Monmouth College), Chelsey McKenna (College of Southern Nevada), Selene Nikaido (University of Central Missouri), Christos Noutsos (SUNY at Old Westbury), Leocadia Paliulis (Bucknell University), Rebecca Seipelt-Thiemann (Middle Tennessee State University), and Melkamu Woldemariam (The College of New Jersey).
Jamboree instructors: Cristina Fernández-Marco (DNA Learning Center - Cold Spring Harbor Laboratory), Marcela Karey Tello-Ruiz (CSHL), and Raj Singh Khangura (Purdue University).
Three additional researchers participated as observers: Jerome Grimplet from the EU-COST grapevine project and Sally Elgin and Wilson Leung from the Genomics Education Partnership (GEP) at Washington University in St. Louis. Observers not only shared their own knowledge, but may help promote future Jamborees within their own faculty training networks.
Over the next year, we will continue supporting faculty attendees via periodic meetings and providing assistance in developing bioinformatics lessons and wet lab resources that can be implemented in the classroom. As we continue to refine the Jamboree approach we will also use the experience gained to help develop similar efforts to improve genomic annotations in other species of the maize pangenome, sorghum, grape, and other important crops.

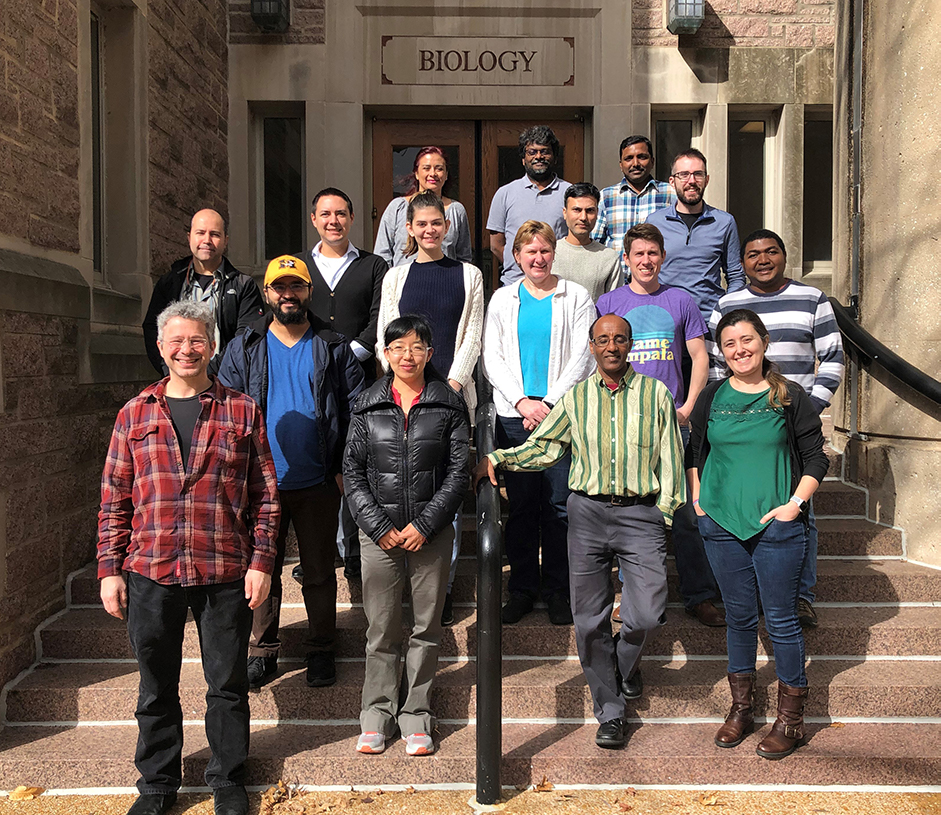
March Jamboree for Researchers
The third Maize Annotation Jamboree—the first designed for researchers—was held on March 13-14, 2019 at the Biology Department of Washington University in St. Louis, Missouri. The objective is for the researchers to establish collaborations with PUI faculty and be part of a larger community curation effort to improve the B73 Zea mays v4 gene models.
Thirteen applicants received scholarships to participate in this two-day event followed by the 2019 Maize Genetics Conference (MGC) Conference.
Jamboree participants: Feseha Abebe-Akele (Texas A&M International University), Michael Jochum (Texas A&M University), Abi Gyawali (University of Missouri, Columbia), Waltram Ravelombola (University of Arkansas, Fayetteville), Shailesh Karre Satyanarayana Guptha (North Carolina State University), Ramesh Dhakal (U of A Rice Research and Extension Center), Ghana Challa (University of Illinois at Urbana-Champaign), Penny Kianian (U Minessota), Ying Hu (U Florida), Patrick Monnahan (U Minessota), Erin Baggs (UC Berkeley); David Carlson (Stony Brook), and Juan Antonio Baeza (Clemson University).
Jamboree instructors: Cristina Fernández-Marco (DNA Learning Center - Cold Spring Harbor Laboratory), Marcela Karey Tello-Ruiz (CSHL), and Joshua Stein (CSHL).
This effort will continue via periodic meetings to discuss progress on the partnerships where maize researchers who participated in this event will serve as consultants for PUI faculty and students participating in CUREs.
Special thanks to Sarah Elgin and Wilson Leung of the Genomics Education Partnership, and Patrick Clark of the Biology Department of Washington University in St. Louis for their support with this event.

2017
Maize Annotation Jamboree
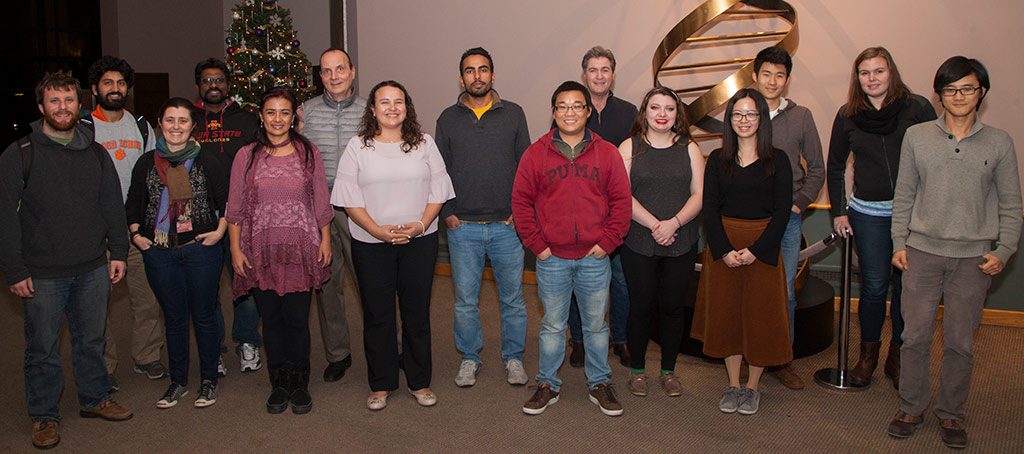
The first genomic annotation jamboree for the current reference Maize B73 (B73, RefGen_V4) was held on December 4-5, 2017 at Cold Spring Harbor Laboratory (CSHL). Sponsored by the NSF-funded MaizeCode (IOS-1445025) and Gramene (IOS-1127112) projects, the jamboree aimed to engage graduate students in the plant research community in the improvement of the Maize gene models. This event was a proof-of-concept for similar future efforts that will help improving annotations in newly sequences maize inbreeds, sorghum, and other important crops.
This event brought together participants from seven US and one international institution (University of Tokyo), and included underrepresented minorities. We also had a participant from University of Toledo, a primarily undergraduate-serving institution (PUI).
Ten graduate students and one postdoctoral applicant were selected to participate in this two-day event: Fei-Man Hsu (University of Tokyo), Hao Wu (Iowa State University), Kokulapalan Wimalanathan (Iowa State University), Junpeng Zhan (University of Arizona), Michelle Stitzer (UC Davis), Pengfei Qiao (Cornell University), Rachel Wasikowski (University of Toledo), Raj Singh Khangura (Purdue University), Sirjan Sapkota (Clemson Univeristy), and Zach Brenton (Clemson Univeristy).
Jamboree instructor: Monica Munoz-Torres, currently a program manager for the development of scientific software at the Translational and Integrative Sciences Lab, at Oregon State University and former project manager of the Apollo project, a web-based genome annotation editor tool designed to support community-based curation. Munoz-Torres gave an introduction on the importance of community-curation efforts and an in-depth demonstration of Apollo’s capabilities. Students were paired groups and tasked with checking the accuracy of 5 distinct maize gene families: PIN, GH3, ABC, TCP and ORC. From these gene families, "suspicious" MAKER-P-generated annotations were identified based on their annotation edit distance (AED) and quality indexes (QI). Working independently on the same set of models, students described their conclusions and approaches and identified ten genes that needed improvements and will be fed back to MaizeGDB curators as updates.
An additional researcher participated as an observer: Uwe Hilgert, director of STEM training at the BIO5 Institute & Cyverse from the University of Arizona.
Jamboree participants continue to annotate their genes of interest and the group will meet periodically to discuss progress and write an official report that we plan on submitting to an educational journal. In addition, Michelle Stitzer presented a summary of the results of this exercise at the Plant and Animal Genome conference on January 15, 2018.

Educational
Resources
Weed to Wonder
The Weed to Wonder website and iPad ebook tells the story of how human ingenuity transformed a common Mexican weed (teosinte) into a modern food wonder (maize). Weed to Wonder shows the continuity of research on corn – from Native American agriculturalists to agricultural breeders, corn geneticists, plant physiologists, and molecular biologists – that culminated in the Maize Genome Sequencing Project. The interactive e-book uses over 150 animations, photographs, illustrations, interviews, and a time-lapse video to provide background on the development of maize, from domestication, hybrid vigor, genome sequencing, and transposons, to genetic modification and biofortification of modern maize. The e-book revolves around footage from Mexico, interviews with prominent scientists, and animations of different approaches to sequencing the maize genome.

Researcher & Graduate Student
Training
Training will prepare faculty from PUIs to analyze MaizeCODE with undergraduate students and provide travel awards for graduate students to attend MaizeCODE training at professional meetings.
Undergraduate Faculty & Student
Outreach
The program will be promote Science, Technology, Engineering and Math (STEM) disciplines by anticipating and encouraging broad participation in primary data analysis by undergraduate and graduate students.


